Tutorial
The following sections provide information to help interpreting MITE entries and creating new submissions.
Overview of the MITE entry page
Creating MITE-compatible reaction SMARTS and SMILES strings
Overview of the MITE entry page
The Navigation Header allows to browse through the MITE entries (the 'left' and 'right' arrow keys on a keyboard can also be used).
The Enzyme Information section provides general information about the tailoring enzyme, including information on Auxiliary Enyzmes (if any are needed for the enzyme to function properly). The Changelog outlines the updates made to this entry.
The Reaction Information column displays the known tailoring reaction of this enzyme. The Reaction SMARTS tab shows the generic reaction as expressed by the reaction SMARTS, while the Reaction Information tab gives additional information.The Example cards below show examples of substrate-product pairs. Multiple tailoring reactions/substrate specificities can be known, and these can be selected clicking on the "Select a reaction" button.

Creating MITE-compatible reaction SMARTS and SMILES strings
Reaction SMARTS and SMILES are the core data in any MITE entry, summarizing the reaction and substrate specificity of the tailoring enzyme. This step-by-step guide shows how to create MITE-compatible SMARTS and SMILES.
What are SMILES, SMARTS, and reaction SMARTS?
SMILES and SMARTS are line notations used to describe chemical structures. SMILES string represents a molecule by specifying its atoms and their connectivity, while a SMARTS string defines a search pattern of atoms and bonds. For example, the SMILES string “C=C” represents ethylene, while the SMARTS string “[C]=[C]” matches both ethylene and propylene (“C=CC”).
Reaction SMARTS extend this concept by modeling chemical reactions, encapsulating the pattern matching of substrates and their conversion to products (“substrate(s)>>product(s)”). This system allows for the modeling of reactions and their application to SMILES strings.
In MITE, reaction SMARTS describe the substrate specificity and reactions of tailoring enzymes, while SMILES are used to represent the actual substrates and products of characterized reactions.
How to create reaction SMARTS and SMILES?
Writing SMILES and reaction SMARTS by hand can be complex and error-prone. Instead, we recommend drawing the chemical structures first and then exporting them as string notations.
Although there are many chemistry drawing tools available, we recommend the free, open-source tool Ketcher, which offers an online demo version and will be used in this tutorial.
Step-by-step guide
Instructions can be also found in this video tutorial.
Example 1: phenol halogenase PltM - specific reaction
In this example, we will characterize the reaction of the enzyme PltM, a phenol halogenase. We will draw the reaction in its easiest form and export the SMILES and reaction SMARTS.
In the publication, PltM is described to halogenate the phloroglucinol substrate, resulting in a mono- or di-halogenated product, with either chloride, bromide, or iodide substituents. Here, we will model the mono-chlorination reaction.
1. In Ketcher, draw phloroglucinol and convert it into its aromatic form by pressing ALT+A. As a rule of thumb, all molecules must be turned into their aromatic (not kekulized) form to be MITE-compatible.

2. Select phloroglucinol by drawing a rectangle (or pressing CTRL+A) and export it as a Daylight SMILES string using the Save Structure menu (pressing CTRL+S). “c1c(O)cc(O)cc1O” is the SMILES string of the substrate.

3. In another Ketcher window, draw the chloro-phloroglucinol product. You can draw the structure from scratch or copy-paste phloroglucinol from the first Ketcher window (CTRL+C/CTRL+V) and add a chloride substituent. Do not forget to aromatize as before with ALT+A. As before, select the molecule by drawing a rectangle (or pressing CTRL+A) and export it as a Daylight SMILES string using the Save Structure menu (pressing CTRL+S). “c1c(O)cc(O)c(Cl)c1O” is the SMILES string of the product.

4. Select the chloro-phloroglucinol and copy it (CTRL+C). Go back to the first window containing phloroglucinol, add a reaction arrow, and paste the chloro-phloroglucinol on the right-hand side of the arrow. This is your reaction.

5. Next, select the reaction (CTRL+A), and select the Reaction Auto Mapping Tool, using the “Discard” mode. This will add paired atom indices to all atoms that are both on the substrate and the product side. Since the chloride atom is only present on the product side, it does not receive an index.

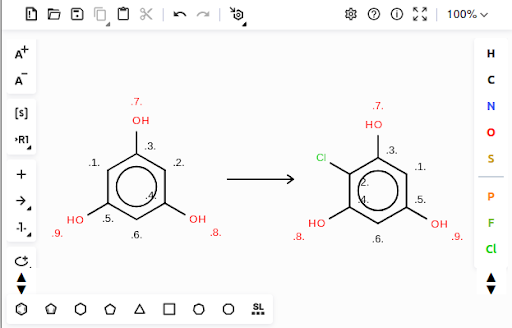
6. Select the reaction again (CTRL+A) and export it as a Daylight SMARTS (CTRL+S). “[#6:1]1:[#6:5](-[#8:9]):[#6:6]:[#6:4](-[#8:8]):[#6:2]:[#6:3]:1-[#8:7]>>[#6:1]1:[#6:5](-[#8:9]):[#6:6]:[#6:4](-[#8:8]):[#6:2](-Cl):[#6:3]:1-[#8:7]” is the reaction SMARTS string. With substrate and product SMILES and reaction SMARTS prepared, the MITE entry can be filled in and submitted.
